To be honest, whenever we talk about medical AI, people usually think of analysing MRI scans or complex genomic data. But they often ignore the most common diagnostic tool: the routine blood test. Currently, clinical hematology mostly just checks if a few biomarkers cross a standard threshold. A massive amount of complex, non-linear information is simply thrown away. Waiting for that incredibly slow, old centrifuge to finish spinning in our engineering building last Friday, I kept thinking about this massive data waste.
In my view, applying deep learning to these everyday blood markers will definitely open a completely new era of preventative medicine [1]. Machine learning models can process heterogeneous datasets to predict disease onset years before symptoms appear. However, fixing the technical bottleneck is only the first step. The destructive “black box” nature and “representational bias” of predictive AI are extremely risky in a real hospital setting [2]. Therefore, we cannot rely on algorithms alone; we must establish a strict “human-in-the-loop” mechanism. In this blog, I will explore how predictive AI is transforming computational hematology, and why we must balance algorithmic accuracy with actual medical humanistic care.
- The Bottleneck: Why Traditional Hematology Needs an AI Intervention
1.1 Defining Computational Hematology in Modern Healthcare
This blog focuses on a specific sub-field within healthcare: Computational Hematology. Traditionally, clinical hematology is about diagnosing blood disorders using microscopes and basic lab tests. However, from an engineering perspective, blood is basically a massive, high-dimensional dataset carrying the continuous health status of our entire body [3]. The other day, while waiting for a 50GB anonymised blood test dataset to slowly download over the notoriously unstable eduroam Wi-Fi in the library, I realised just how data-driven this discipline has become. In simple terms, computational hematology applies machine learning to process these complex, multi-dimensional blood markers.
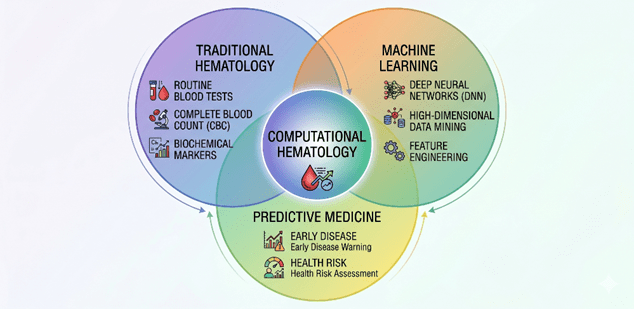
Figure 1: Interdisciplinary Architecture of Computational Hematology.
It shifts the medical focus from treating visible diseases to predicting future health risks using underlying data patterns.
1.2 The Complexity of Current Challenges: Low Utilization & Lagging Prevention
Currently, this field is facing two major systematic problems. The first one is the surprisingly low utilisation of high-dimensional biomarker data. Every single day, millions of routine blood tests are done globally. Yet, the standard clinical workflow remains quite traditional and rigid. Doctors usually just check if a single indicator is out of the “normal threshold”. For example, if a patient’s fasting blood sugar is slightly below the pre-diabetic line, they are often classified as completely healthy. The medical system ignores the complex non-linear connections between dozens of biochemical indicators.
Actually, human biology does not operate on simple step functions. A single marker might look normal on its own, but when you combine its slight deviation with subtle changes in lipid panels and white blood cell counts, it could indicate a serious underlying metabolic issue [4].
On Tuesday evening, whilst rushing to sort through a jumbled CSV file containing heart rate and blood oxygen readings—all required for another assignment—in just an hour at the urging of my group members, it hit me. Staring at those thousands of rows and columns, I realized it is practically impossible for a human doctor to manually calculate these high-dimensional interactions during a brief 10-minute consultation. They simply look for the red flags. Think about the sheer volume of data generated by Complete Blood Counts (CBC). We store them in databases, but rarely use them to track a patient’s longitudinal health trajectory.
This brings us to the second major challenge: the severe lag in preventative medicine. Because our healthcare system heavily relies on these isolated thresholds, traditional blood tests usually only diagnose a condition when the disease has already occurred or when obvious pathological features appear. For instance, by the time certain kidney or liver enzymes cross the clinical threshold, the organ has often already suffered irreversible damage [5]. We are essentially missing the golden window for early intervention. We use blood tests to confirm sickness, rather than to preserve health. This massive waste of predictive data highlights exactly why we must rethink our clinical approach.
1.3 The Rationale for Machine Learning: Decoding High-Dimensional Data
To understand why we urgently need Machine Learning in computational hematology, we first need to look at the fundamental limitations of traditional statistical models. In conventional clinical research, tools like logistic regression or Cox proportional hazards models are the absolute standard. However, these classical methods have a critical weakness when dealing with modern biomedical data: they heavily rely on the assumption that input variables are largely independent.
I still remember looking up the exact mathematical properties of “multicollinearity” on a tech wiki during a late-night study session for my first-year applied statistics module. The encyclopedia article explained how highly correlated predictors can completely destabilise a standard regression model, causing the variance of coefficient estimates to inflate exponentially. Reading that, it struck me that the human body is actually the ultimate multicollinear nightmare. In a routine blood test, biological indicators do not fluctuate in isolation. A subtle drop in red blood cell count is often tightly coupled with specific changes in ferritin levels, liver enzymes, and immune system markers[6]. Traditional biostatistics forces us to manually drop these correlated variables or use dimensionality reduction techniques just to make the math work. Consequently, we deliberately discard potentially vital biological information before the analysis even begins.
This is exactly where Machine Learning (ML) changes the rules. Unlike classical statistics, advanced ML algorithms—such as Deep Neural Networks (DNNs) or tree-based ensemble methods like XGBoost—do not require strict assumptions about data independence or linear boundaries. They are specifically engineered to handle high-dimensional feature spaces without manually filtering out overlapping variables.
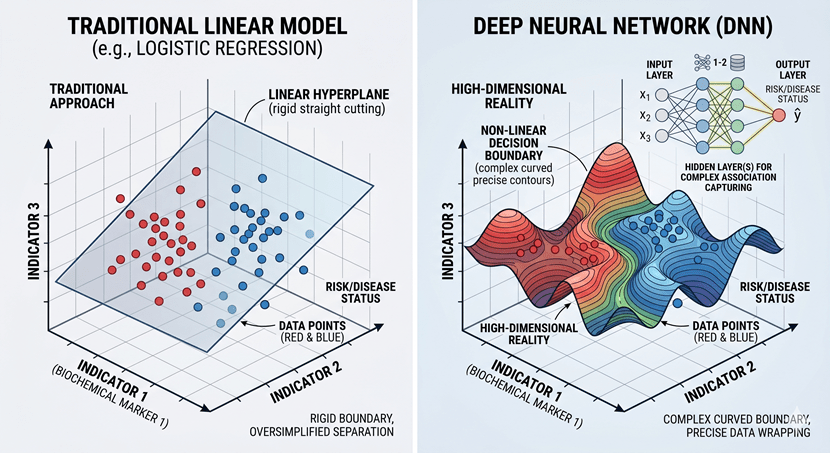
[Figure 2: Decision Boundaries: Traditional Linear Models vs. Deep Neural Networks].
By feeding the algorithm with dozens of raw, unedited biochemical parameters, we allow the model to autonomously learn the complex, non-linear relationships between them. For instance, an ML model can seamlessly detect that a 2% increase in marker A, combined with a 5% decrease in marker B, might be statistically irrelevant on its own but becomes highly predictive of an early-stage cardiovascular event when paired with a specific age group [7].
Additionally, ML is uniquely capable of ingesting massive, heterogeneous datasets. A patient’s health trajectory is rarely captured by a single blood draw taken on a random Tuesday. To build a robust predictive system, we must analyse longitudinal data—repeated tests over months or years—and integrate them with structured electronic health records (EHR). Deep learning architectures, particularly Recurrent Neural Networks (RNNs) or modern attention mechanisms, can track how a patient’s blood matrix evolves over time. They align these sequential fluctuations to identify hidden disease precursors long before any physical symptoms manifest.
In my view, the true justification for adopting predictive AI is not about finding a computationally heavier replacement for doctors. It is about upgrading our diagnostic resolution. Human brains and traditional calculators simply cannot map a 100-dimensional feature space in real-time. ML technologies possess the computational capacity to digest these massive datasets, mining the subtle, hidden patterns of disease from minute biochemical fluctuations.
- Under the Hood: AI/ML Breakthroughs in Blood Matrix Analysis
2.1 The Macro Foundation: Smart Supply Chains and Blood Bank Management
Before we discuss how to extract medical insights from a drop of blood, we must first briefly acknowledge the macro-level foundation: how the blood physically gets to the hospital. Managing the national blood supply chain is essentially a highly complex spatiotemporal routing problem. Blood products, like platelets and red blood cells, have very short shelf lives. Traditional inventory systems struggle with this, which easily causes the coexistence of blood shortages in remote areas and expiration waste in central hospitals. To fix this supply and demand mismatch, researchers are increasingly applying Time-Series forecasting combined with Graph Neural Networks (GNNs) [8]. By modelling blood banks and hospitals as nodes, and transit routes as edges, GNNs can dynamically calculate optimal distribution strategies to prevent expiration waste.
Last Thursday, I was grabbing a quick Tesco meal deal in the freezing London rain between my machine learning lectures. Watching delivery vans completely gridlocked in the traffic, it struck me that optimising blood transport is structurally similar to fixing city logistics. It solves the physical distribution problem.
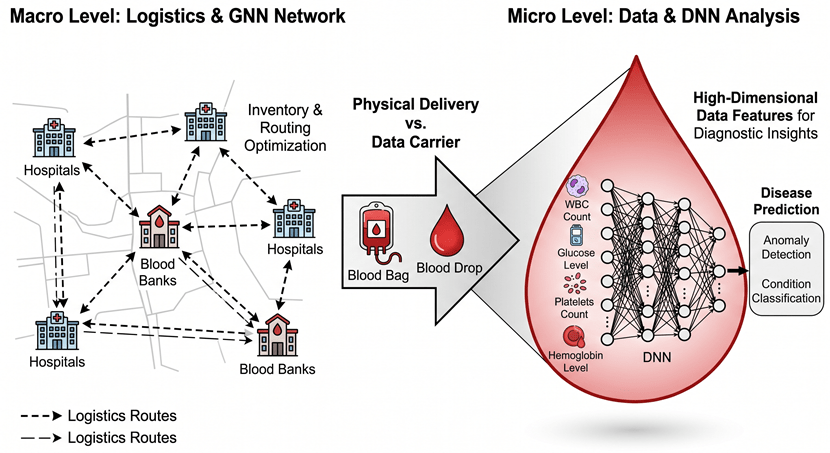
Figure 3: Conceptual Transition from Macro-level Logistics (GNN) to Micro-level Diagnostic Intelligence (DNN).
But here is the catch. While algorithms like GNNs efficiently ensure a bag of blood reaches the emergency room exactly when needed, they treat the blood purely as a consumable commodity. Actually, viewing blood just as a liquid for physical transfusion severely underestimates its potential. Blood samples are also a massive data carrier that contains the health status of our whole body.
Once the macro-level logistics secure the physical supply, we are presented with a biochemical matrix containing a systemic snapshot of the human body. If macro-level AI acts as the physical transportation infrastructure, then micro-level predictive classifiers are the data extraction engines. Therefore, the rest of this blog will shift the focus away from the macro supply chain. Instead, we will concentrate entirely on how deep learning models dig into these routine blood datasets to decode hidden disease patterns.
2.2 Micro-Diagnostics: Predictive Classifiers and Deep Neural Networks
2.2.1 Technical Principles
Now that we have established the macro-level logistics as our background, we can focus on the core technical anchor of this blog: the micro-level predictive classifiers. Specifically, we need to understand exactly how machine learning models process high-dimensional blood biomarkers, such as Complete Blood Counts (CBC) and lipid metabolites.
To explain this, I often use a specific analogy during our lab seminars. Think of the human body’s blood matrix as a massive symphony orchestra. A traditional diagnostic approach is like listening to just the first violin (e.g., fasting glucose) and the cello (e.g., LDL cholesterol). If they are playing the right notes, the doctor assumes the whole piece is fine. But a chronic disease in its early stage is rarely a single wrong note; it is a subtle dissonance spread across the entire orchestra. Human ears, or traditional linear models, cannot easily detect this distributed disharmony.
This is where algorithms like Gradient Boosting Machines (specifically XGBoost) and Deep Neural Networks (DNNs) come in. They are designed to listen to the entire symphony at once.
Let us look at XGBoost first. In my view, while deep learning gets most of the media hype, tree-based ensemble methods are actually often more robust for tabular clinical data like blood tests[9]. XGBoost works by building sequential decision trees. Instead of trying to find one perfect equation to separate sick patients from healthy ones, it trains a simple tree, calculates the residual errors (the patients it misclassified), and then builds the next tree specifically to correct those errors. When processing a CBC panel, XGBoost naturally captures complex, conditional interactions. For example, it might learn that a slightly elevated white blood cell count is only a risk factor if the patient’s triglyceride level is also above a certain threshold and their age is over 45. It maps these non-linear clinical rules without needing us to manually specify them.
On the other hand, Deep Neural Networks (DNNs) take a different mathematical approach: representation learning. A typical CBC and metabolic panel might contain 30 to 50 raw variables. A DNN feeds these raw numbers into an input layer, passing them through multiple hidden layers using non-linear activation functions, such as ReLU.
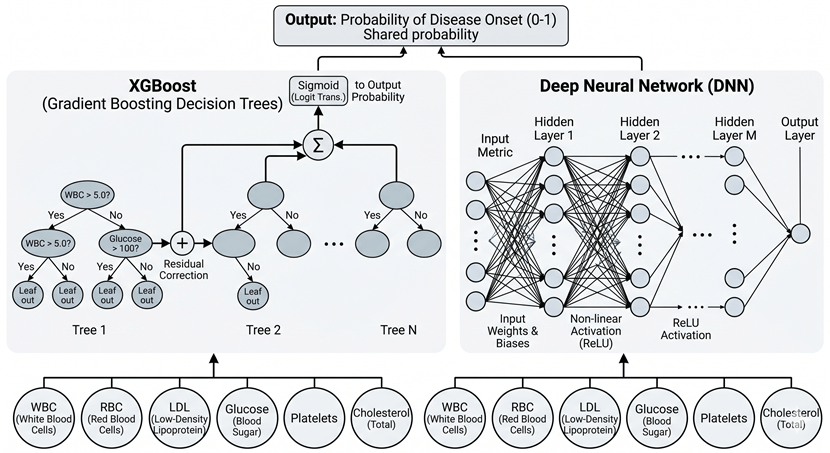
Figure 4: Comparison of Architectural Paradigms between XGBoost and Deep Neural Networks for Blood Biomarker Analysis.
During this forward pass, the DNN transforms the raw, noisy blood markers into a highly abstract, lower-dimensional latent space. Basically, the first hidden layer might detect basic correlations between red blood cells and iron levels, while the deeper layers combine these patterns to form a high-level representation of “systemic inflammation” or “metabolic stress” [10]. Through backpropagation, the network continuously adjusts its internal weights to minimise the prediction error.
To be honest, both XGBoost and DNNs achieve the same fundamental goal here. They discard the outdated “single-variable threshold” method and instead construct a multidimensional decision boundary. By mathematically digesting the interactions among dozens of blood markers simultaneously, they can spot the biological precursors of a disease long before it causes any physical symptoms.
2.2.2 Technical implementation challenges
While the mathematical architecture of Deep Neural Networks (DNNs) and XGBoost sounds incredibly powerful on paper, deploying them in real-world computational hematology is a completely different story. To be honest, if you feed raw, unedited hospital data directly into these algorithms, they will almost certainly fail. The core problem lies in the inherent nature of biomedical data, which is constantly plagued by high noise and systematic bias.
First, let us consider biological noise. Blood is not a static substance; it is a highly dynamic fluid. A patient’s biomarker levels fluctuate significantly based on what they ate for breakfast, their acute stress levels, or even the time of day the blood was drawn. This creates a massive amount of intra-patient variance [11].
Second, and perhaps more problematic for machine learning engineers, is the issue of systematic bias, frequently referred to in bioinformatics as ‘batch effects’. Different hospitals and clinics do not use the same analytical instruments. A central hospital in London might use a Sysmex hematology analyzer, while a local clinic in Manchester relies on a Beckman Coulter machine. These instruments utilize varying calibration standards, optical sensors, and chemical reagents. Therefore, a hemoglobin reading of 130 g/L from machine A is not mathematically identical to a 130 g/L reading from machine B [12]. If we are not careful, a highly capable deep learning model might just learn to classify which hospital the patient went to, rather than predicting their actual cardiovascular risk.
To understand this in a simpler way, think of it like training an AI to recognize a specific melody. The hidden disease pattern is the melody we want. However, the audio data we receive is recorded at a bustling train station (which represents biological noise), using dozens of completely different, poorly calibrated microphones (which represents instrument bias). If we just feed the raw audio to the ML model, it will likely memorize the background train announcements or the specific microphone distortion instead of the song itself.
To overcome these implementation hurdles, our systems heavily rely on rigorous Feature Engineering and advanced Representation Learning. Simple techniques like standard z-score normalization are often insufficient here, because the underlying patient demographics at a specialist oncology hospital are fundamentally different from those at a general clinic. Instead, we must use advanced imputation methods like MICE (Multiple Imputation by Chained Equations) to logically handle the inevitable missing values, and apply quantile normalization to align the distributions across different laboratories.
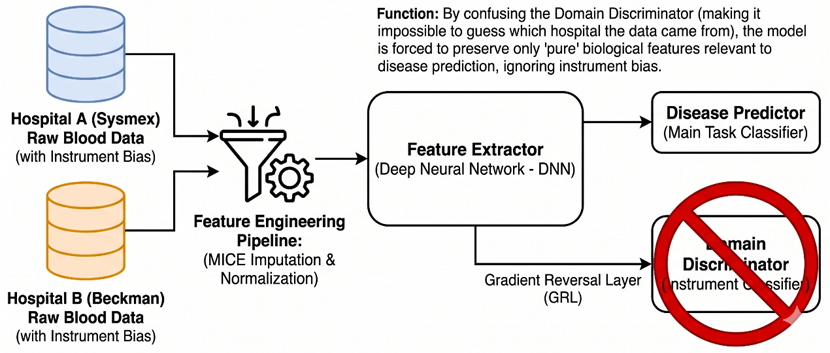
Figure 5: Architecture of the Domain Adversarial Neural Network (DANN) for Instrument Bias Removal and Biological Feature Extraction.
Furthermore, we implement Representation Learning techniques directly within our neural network architectures. For instance, we can utilize Domain-Adversarial Neural Networks (DANN). The objective here is to force the hidden layers to extract data features that are highly predictive of the disease, but entirely invariant to the source instrument. We are basically forcing the network to mathematically strip away the “microphone distortion” and only listen to the biological “melody” [13].
In my view, managing these severe data imperfections is where the real engineering challenge lies. A predictive classifier is ultimately only as robust as the data representation it learns from.
2.2.3 Breakthroughs in application scenarios
Once we have effectively managed the biological noise and instrument biases, the true potential of these predictive classifiers becomes apparent in real clinical scenarios. The most significant breakthrough is shifting the diagnostic timeline. We are moving from confirming a disease that has already occurred to calculating the probability of a disease occurring in the next 3 to 5 years.
To illustrate this, think of a patient’s health as a large, complex jigsaw puzzle. A traditional blood test is like looking at five or six disconnected puzzle pieces and trying to guess the final picture. A machine learning model, however, looks at all 50 pieces simultaneously, recognises the subtle patterns connecting them, and predicts what the completed picture will look like before the remaining pieces are even placed on the table.
Let us look at cardiovascular disease (CVD) prediction. Traditionally, doctors rely heavily on lipid panels—specifically LDL cholesterol—to assess heart attack risks. However, many patients suffer heart attacks despite having “normal” cholesterol levels. A trained Deep Neural Network looks much deeper. It might detect a complex interaction: a patient has high-normal LDL, but also shows a very slight decrease in mean corpuscular volume (MCV) and a marginal elevation in specific inflammatory markers like CRP. While any human doctor would dismiss these tiny fluctuations as normal variance, the algorithm identifies this specific multi-variable signature as a high-risk precursor to arterial plaque rupture within the next 3 years [14].
A similar breakthrough is happening with Type 2 Diabetes. The clinical standard is to monitor fasting glucose or HbA1c. But by the time these metrics cross the diagnostic threshold, pancreatic beta-cell function is often already severely compromised. Using XGBoost, researchers have successfully predicted the onset of diabetes up to 5 years in advance [15]. The model achieves this by mapping the non-linear relationships between liver enzymes, triglyceride ratios, and subtle shifts in the white blood cell differential—a pattern that indicates systemic insulin resistance long before the blood sugar actually spikes.
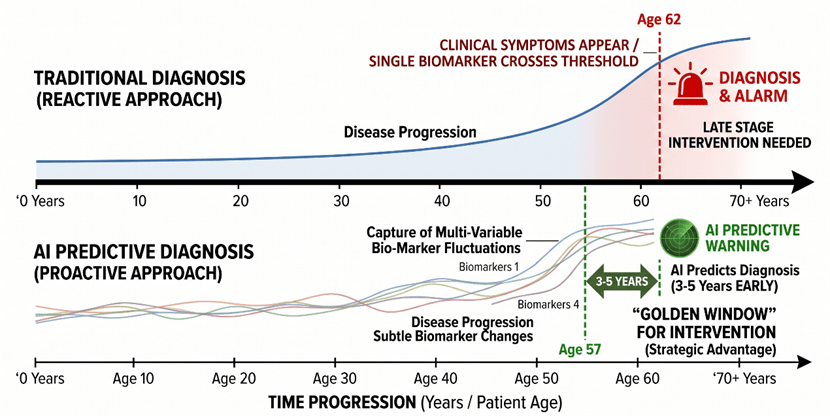
Figure 6: Timeline Comparison of Traditional vs. AI-Predictive Diagnostics and the “Golden Window” for Intervention.
Finally, the most acute application is sepsis prediction in intensive care units. Sepsis is a rapid, life-threatening inflammatory response. Every hour of delayed treatment drastically increases the mortality rate. Traditional scoring systems, like SOFA, are often too slow. DNNs, continuously fed with real-time blood test results and vital signs, can identify the hidden “sepsis signature”—a cascading failure pattern among immune cells and coagulation factors. They can predict septic shock 4 to 12 hours before clinical symptoms appear, giving doctors a critical window to administer antibiotics [16].
In my view, these applications prove that predictive ML is not just a statistical upgrade; it fundamentally reshapes preventative medicine. It forces us to redefine what “healthy” means in a clinical setting.
3. The Ethical Labyrinth: Governance and Humanistic Care in Predictive AI
3.1 Representational Bias and the Threat to Health Equity
Up to this point, we have focused on the technical performance of predictive AI. But technical accuracy in a controlled lab does not automatically equal clinical safety in the real world. To be honest, the biggest threat to computational hematology right now is not the algorithm architecture itself, but the raw data we feed it. This brings us to a critical ethical issue: representational bias and health equity.
In machine learning, we often naturally assume that a massive dataset is an objective reflection of reality. Actually, biomedical data is deeply shaped by socio-economic factors. The large-scale electronic health record (EHR) datasets used to train these deep neural networks usually come from top-tier research hospitals and biobanks. This means the training data is heavily skewed towards specific socio-economic classes or ethnic groups who have regular access to high-quality, preventative healthcare. Marginalized groups, low-income populations, or racial minorities—who might only visit the hospital during severe emergencies—are systematically underrepresented in the training matrix [17].
This lack of representation becomes extremely dangerous because human blood baselines are not universally identical. There are natural, healthy biological variations in blood markers across different races, genders, and environments. If a model is trained on a homogeneous dataset, it will mathematically establish a narrow, privileged definition of “normal” that simply does not apply to everyone.
Let me share a real and widely discussed clinical case that perfectly highlights this danger. In hematology, there is a well-documented condition known as Benign Ethnic Neutropenia (BEN). Many healthy individuals of African and Middle Eastern descent naturally have lower absolute neutrophil counts (a vital type of white blood cell) compared to Caucasian populations. This lower count is completely normal and healthy for their genetics. However, if we deploy a predictive deep learning model trained predominantly on white European blood profiles, the algorithm will mathematically interpret this naturally lower neutrophil count as an anomaly. It might flag it as a severe immune deficiency or an early sign of bone marrow failure [18].
The clinical consequences of this are severe. For marginalized groups, the blind application of such uncalibrated models leads to two direct outcomes: high false positives and high false negatives. In the BEN example, the model generates a dangerous false positive. It could potentially subject a perfectly healthy Black patient to unnecessary anxiety, invasive bone marrow biopsies, or even cause doctors to delay crucial chemotherapy treatments because the algorithm mistakenly insists their immune system is too weak to handle it. Conversely, a false negative occurs when a model fails to detect an actual disease because a minority patient’s abnormal biomarker shift happens to look like the “healthy” baseline of the majority training group.
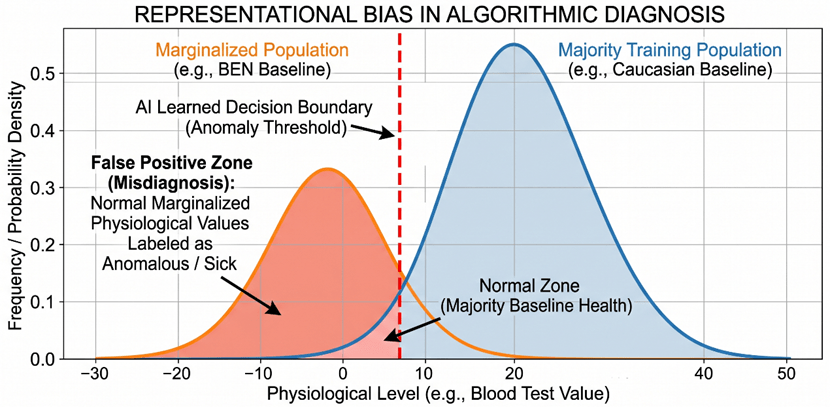
Figure 7: Representational Bias and the Misalignment of Algorithmic Decision Boundaries between Majority and Marginalized Populations.
In my view, this is exactly why we cannot treat AI alignment purely as a software debugging task. During our recent seminars on LLM alignment, we discussed the concept of “pluralistic alignment”—the idea that AI must align with diverse cultural values. In computational hematology, pluralistic alignment means our algorithms must mathematically respect biological diversity. We cannot blindly deploy a single predictive model across diverse populations without validating its cross-demographic performance.
If we ignore representational bias, these predictive classifiers will not democratise healthcare. Instead, they will act as a technological amplifier, quietly widening the existing medical inequality gap under the guise of objective mathematics. Engineering an accurate model is only half of our job; ensuring that the model serves all populations fairly and safely is the real engineering challenge.
3.2 The Black Box Problem and the Explainability Crisis
Beyond biased data, we must face another fundamental issue with Deep Neural Networks (DNNs): the “Black Box” problem. To be honest, as engineering students, we often obsess over optimising metrics like the Area Under the Curve (AUC) or predictive accuracy. But in clinical settings, high mathematical accuracy alone is not enough. If an algorithm predicts a patient has a 90% chance of developing acute leukemia in the next three years based on a routine blood test, the immediate question from both the doctor and the patient will be: “Why?”
Because of the complex, non-linear hidden layers we discussed earlier, a standard DNN cannot answer this. It gives a probability, but it cannot output a medical reasoning path. This lack of transparency directly conflicts with current legal frameworks. For instance, under the European Union’s GDPR, specifically Article 22, individuals have the right to obtain “meaningful information about the logic involved” in automated decision-making [19]. A black-box output simply fails this legal requirement.
To understand how dangerous this opacity can be, we can look at a classic real-world failure in medical machine learning. Researchers once trained a neural network to predict pneumonia survival rates to decide which patients should be admitted to the hospital. The model achieved fantastic accuracy. However, because it was a black box, the developers did not initially realise that the model had learned a lethal rule: it predicted that patients with asthma had a lower risk of dying from pneumonia. Actually, this happened because asthma patients in the training data were automatically sent to the Intensive Care Unit (ICU), and this aggressive treatment lowered their mortality rate in the records [20]. If doctors had blindly trusted this highly accurate black box without asking “why”, they would have sent high-risk asthma patients home to die.
This brings us to a tough debate. In life-or-death medical scenarios, does a purely “highly accurate” model have clinical legitimacy if it lacks explainability? In my view, the answer is no. Doctors hold the ultimate legal and moral responsibility for patient care. They cannot ethically prescribe a heavy, toxic chemotherapy regimen based on an algorithm’s unexplainable mathematical intuition.
Therefore, integrating eXplainable AI (XAI) techniques, such as SHAP (SHapley Additive exPlanations) or LIME, is no longer just an optional engineering upgrade [21]. These tools approximate the black box with interpretable models, highlighting exactly which specific blood markers drove the prediction. Without this transparency, we cannot build clinical trust. A predictive AI in hematology must act as an interpretable diagnostic co-pilot, not an unquestionable oracle.
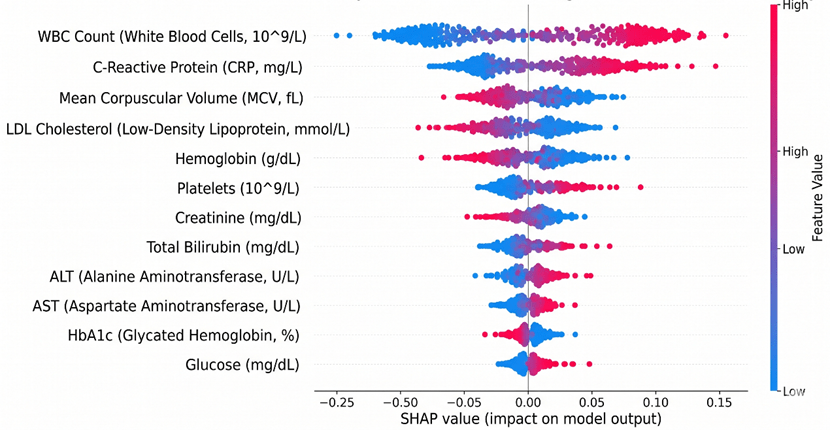
Figure 8: SHAP plot illustrating the contribution of key blood biomarkers to the disease prediction model.
3.3 Psychological Impact and the Necessity of the “Human-in-the-Loop”
Even if we completely solve the technical issues of representational bias and the black box, we still face a purely human problem. To be honest, mathematical accuracy does not automatically translate into good medical practice. A predictive blood matrix model essentially acts as a medical oracle. It tells you what might happen years before it actually does. But an algorithm has zero empathy. If an XGBoost model predicts that a healthy 30-year-old blood donor has an 85% probability of developing severe heart failure within the next five years, how exactly do we deliver this news?
Directly pushing these terrifying prediction scores to a patient’s smartphone or email is highly irresponsible. The psychological impact of predictive medicine is enormous. Knowing you have a high probability of getting a severe disease in the future creates a unique type of chronic anxiety [22]. The algorithm only provides a cold percentage, but the patient is left alone with the existential dread of that number.
We have actually seen the devastating consequences of removing the human buffer in automated medical communication. I recall a very controversial issue right here in the UK recently concerning the NHS App. Due to automated record-sharing policies, many patients received push notifications and read their own biopsy results—confirming terminal cancer—directly on their phones before their General Practitioner (GP) even had the chance to call them. Patients were left sitting alone in their living rooms, staring at a screen, completely panicking because there was no medical professional there to explain the context, answer their questions, or offer a treatment plan. If we allow predictive AI to automatically send future disease risks to blood donors, we are going to scale up that exact same psychological trauma.
This is why we must strongly advocate for a mandatory “Human-in-the-loop” (HITL) framework in computational hematology. The AI should strictly serve as a diagnostic co-pilot, not the final decision-maker or communicator. The output of the deep neural network—along with its SHAP value explanations—must go to a human doctor first.
In this framework, the doctor’s role shifts from manually computing raw blood data to providing essential humanistic care. The doctor evaluates the AI’s prediction, considers the patient’s current mental state, and then delivers the news with genuine empathy. More importantly, they provide an actionable preventative strategy. An algorithm can calculate a risk score from a CBC panel, but only a human doctor can look a patient in the eye and say, “The system caught a warning sign early, but here is exactly what we are going to do together to prevent it.”
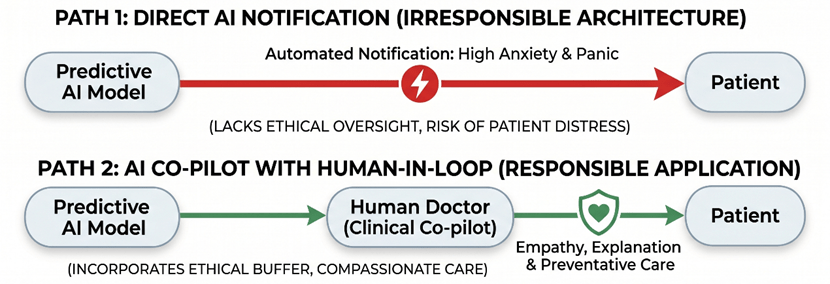
Figure 9: Comparison of Healthcare AI Deployment Architectures: Direct Automated Notification vs. Human-in-the-Loop (HITL) Ethical Buffer.
In my view, predictive machine learning is undoubtedly the greatest technical breakthrough in modern hematology. It unlocks the massive hidden value of routine blood tests. However, we must remember that medicine is fundamentally a human-centric discipline. The more powerful and complex our predictive algorithms become, the more urgently we need human doctors to act as the ethical buffer. True medical progress is not just about engineering the most accurate neural network; it only happens when algorithmic accuracy is matched with actual humanistic care.
Conclusion: The Future of Hematology is Human
To bring this discussion together, predictive machine learning has fundamentally rewritten the rules of what we can achieve with a simple drop of blood. By breaking free from the limits of traditional single-variable thresholds, we have successfully unlocked the massive predictive potential hidden inside routine clinical data. Algorithms like Deep Neural Networks are driving a true revolution in computational hematology. They allow us to shift from reactive disease confirmation to proactive prevention, buying patients critical years for early intervention.
However, pushing mathematical boundaries is only half the battle. As we have seen, deploying these highly capable models introduces dangerous blind spots. The opacity of black-box architectures and the embedded representational bias in training data can easily translate into real-world medical inequality.
To be honest, medicine is never just pure data science; it is fundamentally a discipline of human care. The more powerful and mathematically unexplainable our predictive AI becomes, the more urgently we need human doctors to stay in the loop. Actually, their role is not diminishing—it is elevating. We rely on medical professionals to act as the ultimate ethical buffer and the final decision-makers. They are the only ones who can filter out algorithmic bias and translate cold risk percentages into empathetic, actionable medical advice. Ultimately, the dividends of AI innovation in healthcare will only be realized if we firmly anchor our engineering efforts on the foundations of fairness, clinical trust, and humanistic care.
Reference
[1] Alimohammadi, A. (2024). CAN: a deep learning approach to diagnose chronic kidney disease.
[2] Alderman, J. E., Palmer, J., Laws, E., McCradden, M. D., Ordish, J., Ghassemi, M., … & Liu, X. (2025). Tackling algorithmic bias and promoting transparency in health datasets: the STANDING Together consensus recommendations. The Lancet Digital Health, 7(1), e64-e88.
[3] Nazha, A., Elemento, O., Ahuja, S., Lam, B., Miles, M., Shouval, R., … & Haferlach, T. (2025). Artificial intelligence in hematology. Blood.
[4] Bennett, L., Mostafa, M., Hammersley, R., Purssell, H., Patel, M., Street, O., … & ID-LIVER Consortium. (2023). Using a machine learning model to risk stratify for the presence of significant liver disease in a primary care population. Journal of Medical Artificial Intelligence, 6.
[5] Deepa, D. R., Sadu, V. B., & Sivasamy, D. A. (2024). Early prediction of cardiovascular disease using machine learning: Unveiling risk factors from health records. AIP Advances, 14(3).
[6] Wang, X., Zhang, D., Lu, L., Meng, S., Li, Y., Zhang, R., … & Gao, R. (2025). Development and validation of an explainable machine learning model for predicting the risk of sleep disorders in older adults with multimorbidity: a cross-sectional study. Frontiers in Public Health, 13, 1619406.
[7] Lin, T. H., Chung, H. Y., Jian, M. J., Chang, C. K., Lin, H. H., Yen, C. T., … & Shang, H. S. (2025). AI-driven innovations for early sepsis detection by combining predictive accuracy with blood count analysis in an emergency setting: retrospective study. Journal of Medical Internet Research, 27, e56155.
[8] Thakur, S. K., Sinha, A. K., Negi, D. K., & Singh, S. (2024). Forecasting demand for blood products: Towards inventory management of a perishable product. Bioinformation, 20(1), 20.
[9] Grinsztajn, L., Oyallon, E., & Varoquaux, G. (2022). Why do tree-based models still outperform deep learning on typical tabular data?. Advances in neural information processing systems, 35, 507-520.
[10] Singh, V., Chaganti, S., Siebert, M., Rajesh, S., Puiu, A., Gopalan, R., … & Kamen, A. (2025). Deep learning-based identification of patients at increased risk of cancer using routine laboratory markers. Scientific Reports, 15(1), 12661.
[11] Foy, B. H., Petherbridge, R., Roth, M. T., Zhang, C., De Souza, D. C., Mow, C., … & Higgins, J. M. (2025). Haematological setpoints are a stable and patient-specific deep phenotype. Nature, 637(8045), 430-438.
[12] Sáez, C., Gutiérrez-Sacristán, A., Kohane, I., García-Gómez, J. M., & Avillach, P. (2020). EHRtemporalVariability: delineating temporal data-set shifts in electronic health records. Gigascience, 9(8), giaa079.
[13] Ji, Y., Gao, Y., Bao, R., Li, Q., Liu, D., Sun, Y., & Ye, Y. (2023, June). Prediction of covid-19 patients’ emergency room revisit using multi-source transfer learning. In 2023 IEEE 11th International Conference on Healthcare Informatics (ICHI) (pp. 138-144). IEEE.
[14] Tasmurzayev, N., Amangeldy, B., Baigarayeva, Z., Boltaboyeva, A., Imanbek, B., Maeda-Nishino, N., … & Baidauletova, A. (2025). Enhancing Cardiovascular Disease Classification with Routine Blood Tests Using an Explainable AI Approach. Algorithms, 18(11), 708.
[15] Khattab, A., Chen, S. F., Sadaei, H. J., Wineinger, N. E., & Torkamani, A. (2026). Development and validation of a two-stage machine learning model for personalised type 2 diabetes screening in the All of Us Research Program and UK Biobank. BMJ open, 16(1), e103171.
[16] Futoma, J., Hariharan, S., & Heller, K. (2017, July). Learning to detect sepsis with a multitask Gaussian process RNN classifier. In International conference on machine learning (pp. 1174-1182). PMLR.
[17] Gianfrancesco, M. A., Tamang, S., Yazdany, J., & Schmajuk, G. (2018). Potential biases in machine learning algorithms using electronic health record data. JAMA internal medicine, 178(11), 1544-1547.
[18] Merz, L. E., & Achebe, M. (2021). When non-Whiteness becomes a condition. Blood, The Journal of the American Society of Hematology, 137(1), 13-15.
[19] Van Kolfschooten, H. B. (2024). A health-conformant reading of the GDPR’s right not to be subject to automated decision-making. Medical law review, 32(3), 373-391.
[20] Caruana, R., Lou, Y., Gehrke, J., Koch, P., Sturm, M., & Elhadad, N. (2015, August). Intelligible models for healthcare: Predicting pneumonia risk and hospital 30-day readmission. In Proceedings of the 21th ACM SIGKDD international conference on knowledge discovery and data mining (pp. 1721-1730).
[21] Agrawal, R., Gupta, T., Gupta, S., Chauhan, S., Patel, P., & Hamdare, S. (2025). Fostering trust and interpretability: integrating explainable AI (XAI) with machine learning for enhanced disease prediction and decision transparency. Diagnostic Pathology, 20(1), 105.
[22] Iacono, D. (2025). A Double-Edged Sword: Ethical and Psychological Implications of APOE Genotype Disclosure Across the Lifespan. Neuroethics, 18(3), 51.
留下评论